- Home
- Information
- Miscellaneous
- Bayes
Bayes
Introduction
How does MCMC work?
Let us first consider the starting point for our further considerations, Bayes' theorem:
\[ P( \vec{\theta} | \vec{x}) = \frac{ P( \vec{x} | \vec{\theta}) \cdot P( \vec{\theta})} {P(\vec{x})}\]
The expression \(P(\vec{\theta}│\vec{x})\) describes the probability for the model parameters \(\vec{\theta}\) given the data \(\vec{x}\) and is referred to as the posterior.
In most applications, we are confronted with the following question: We have a model or hypothesis that can be described by the parameters \(\vec{\theta}\) . These are usually unknown. And we have the data \(\vec{x}\), which in most cases has been determined by one or more measurements.
Bayes' theorem shows a way in which the probability distribution \(P( \vec{\theta} | \vec{x}) \) can be determined from other probability distributions.
First, there is the so-called likelihood \(P( \vec{x} | \vec{\theta})\). This describes the probability distribution for the model or hypothesis (with the parameters \(\vec{\theta} \) to be determined, which we assume best describe the distribution of the data \( \vec{x} \)).
The prior \( P(\vec{\theta}) \) describes the probability distribution for our assumptions about the individual parameters \( \vec{\theta} \), which we have without knowledge of the (new) data \( \vec{x} \). The prior therefore reflects our prior knowledge or our assumptions about the parameters. Likelihood and prior are multiplied together and form the numerator of the Bayes’ theorem.
The expression in the denominator, known as the evidence \( P(\vec{x}) \), describes the probability that the data \( \vec{x} \) is described by our chosen model or hypothesis. In principle, this can be calculated by integrating over all possible parameter values \( \vec{\theta} \) .
\[ P( \vec{x}) = \int \limits_{\vec{\theta}} P(\vec{x} | \vec{\theta}) \cdot d\vec{\theta} \]
As already mentioned, it can in principle be calculated in this way. Even for the simplest models, the integral cannot usually be calculated in closed form.
A suitable approach to tackle this problem is to use Markov Chain Monte Carlo (MCMC) algorithms. These are based on the creation of a Markov chain, which is then used for Monte Carlo approximations. At first glance, this sounds quite complicated. We will show with a few examples that this is not the case, at least for simple applications.
Posterior for normally distributed data
Let's start with a simple example. Given measurement data \(\vec{x}\) that we know is normally distributed around the value 0. The standard deviation of the normal distribution is 1.
The following Python code (for use in a Jupyter notebook) simulates our measurement data.
# package provides support for matplotlib to display figures directly inline in the Jupyter notebook
%matplotlib inline
# import modules
import numpy as np
import scipy as sp
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import norm
sns.set_style('whitegrid') # control the general style of the plots: dict, or one of {darkgrid, whitegrid, dark, white, ticks}
sns.set_context('notebook') # control the scaling of plot elements: dict, or one of {paper, notebook, talk, poster}
np.random.seed(123) # initialize the random number generator
numdata = 25 # number of datapoints
data = np.random.randn(numdata)
ax = plt.subplot()
sns.histplot(data, kde=False, ax=ax) == ax.set(title='Histogram of observed data', xlabel='interval', ylabel='# observations');
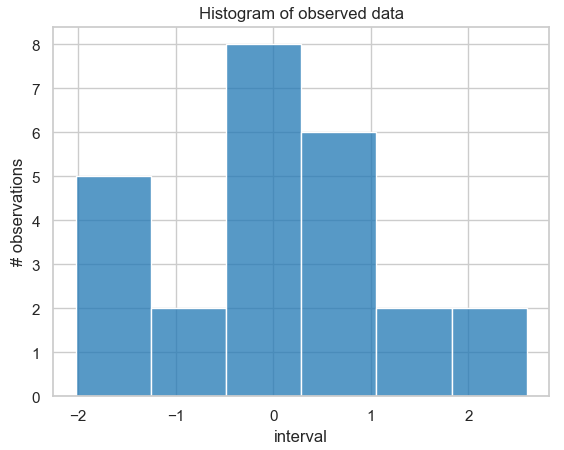
Histogram of the probability distribution of 25 simulated measurement data points that are normally distributed around the value 0 with a standard deviation of 1.
Since we know that we used a normal distribution with a mean of \(\mu=0\) and a standard deviation of 1 for the simulation, we can also determine the solution we are looking for analytically in this exceptional case and use it later for comparison purposes. The reason for this is that for a normally distributed likelihood with known standard deviation, the normally distributed prior is conjugate (i.e., the posterior has the same distribution as the prior). Thus, we know that the posterior with mean μ is also normally distributed.
Here is the Python code for calculating the analytical solution:
# function for calculating posterior analytically
def calc_posterior_analytical(data, x, mu_0, sigma_0):
sigma = 1. # standard deviation is fixed!
n = len(data) # number of data
mu_post = (mu_0 / sigma_0**2 + data.sum() / sigma**2) / (1. / sigma_0**2 + n / sigma**2)
sigma_post = (1. / sigma_0**2 + n / sigma**2)**-1
return norm(mu_post, np.sqrt(sigma_post)).pdf(x)
ax = plt.subplot()
x = np.linspace(-1, 1, 500)
posterior_analytical = calc_posterior_analytical(data, x, 0., 1.)
ax.plot(x, posterior_analytical)
ax.set(xlabel='mu', ylabel='belief', title='Analytical posterior');
sns.despine()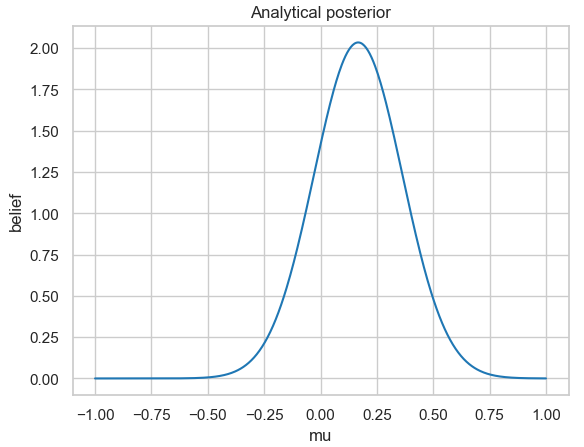
Analytically calculated posterior probability distribution for the 25 simulated normally distributed measurement data with a fixed standard deviation of 1.
If you look closely at the figure, you will see that for the analytical solution, the posterior distribution does not have a mean of \(\mu = 0\). This is because we only determined a small number of measurement data randomly distributed from a normal distribution.
For further study:
Run the code with higher values for the number of simulated measurement data in a Jupyter Notebook and observe how the analytical solution behaves.
After these preparatory steps, let's now turn to our actual task: we want to determine the posterior distribution \( P(\vec{\theta} | \vec{x}) \) from the Bayes’ theorem. We already have the simulated measurement data \( \vec{x} \) from above.
Next, we need the likelihood function \( P( \vec{x} | \vec{\theta}) \). For our simple example, let's assume that the model is a normal distribution.
Of course, you will now say that this is logical, since we simulated our measurement data from a normal distribution. So why should we use a different model now? That's right! But we only used the simulation to generate measurement data. Now let's just “forget” how we generated it. Our starting points (assumptions) for the next steps are then:
- We have a number of n measurement data.
- Each measured value is equal to the initial value (in this case, the value of the normal distribution), i.e., no additional properties such as detector efficiencies or distance dependencies, etc., need to be taken into account (more on this later).
- We assume a normal distribution as a model (we could also assume a different distribution, e.g., a Poisson distribution, a Student's distribution, etc. However, if we look at the probability distribution, it could also be a normal distribution. What happens if we choose the “wrong” model? We will deal with that later).
It follows that the likelihood function is also a normal distribution! It has two parameters, the mean \(\mu\) and the standard deviation \(\sigma\), which we set to 1 in our example.
Our goal is to determine the posterior \( P(\vec{\mu} | \vec{x}) \), with \((\vec{\mu} = \mu) \), since we only have one parameter in our model, the mean \(\mu\) (\(\sigma\) has a fixed value). We can write the model in mathematical notation as follows:
\[ \mu \sim ( \vec{x} | \mu, 1) \]
However, we still have the problem that we do not know the evidence \( P(\vec{x}) \) and normally cannot calculate it analytically. This is where MCMC sampling, the Markov Chain Monte Carlo sampling method, comes into play. By applying it, we do not have to calculate the integral at all!
The "trick" is simply that the algorithm does not calculate the posterior directly, but uses the ratio of the two posteriors for the values \(\mu\) and \(\mu_0\) (note: If you don't believe it, calculate the ratio for the corresponding normalized posteriors and check the result with the non-normalized posteriors; in this simple example, it's not that difficult).
The evidence is responsible for normalizing the solution, the posterior. The integral is determined for the given measured values \(\vec{x}\) for all values of \(\mu\), i.e., the value of the integral is not influenced by which μ is currently used in the Bayes’ theorem.
\[ P( \vec{x}) = \int \limits_{\vec{\theta}} P(\vec{x} | \vec{\theta}) \cdot d\vec{\theta} = \int \limits_{\mu} P(\vec{x} | \mu) \cdot d\mu \]
In our example with the (single) parameter \(\mu\), we set the two posteriors for this
\[ P( \mu | \vec{x}) = \frac{ P( \vec{x} | \mu ) \cdot P( \mu )} {P(\vec{x})}\]
and
\[ P( \mu_0 | \vec{x}) = \frac{ P( \vec{x} | \mu_0 ) \cdot P( \mu_0 )} {P(\vec{x})}\]
in relation as follows (\(\mu\) and \(\mu_0\) are describing two (different) values of parameter \(\mu\)):
\[ \frac { \frac{ P( \vec{x} | \mu ) \cdot P( \mu )} {P(\vec{x})} }
{ \frac{ P( \vec{x} | \mu_0 ) \cdot P( \mu_0 )} {P(\vec{x})} } =
\frac{ P( \vec{x} | \mu ) \cdot P( \mu ) }
{ P( \vec{x} | \mu_0 ) \cdot P( \mu_0 )} \]
Then \(P(\vec{x})\) cancels out (for the expression of the normalized posterior, we set \(\mu_0\), for the non-normalized one, we set \(\mu\))!
How is all this implemented in the MCMC algorithm (using the Metropolis sampler)?
We start with a random initial value for the quantity we are looking for, the mean \(\mu\).
mu_current = 1. # starting parameter - can be randomly choosenThis starting value is the mean of a normal distribution with fixed standard deviation \(\sigma\).
proposal_width=.5 # standarddeviation of normal distributionNote that this normal distribution has nothing to do with the normal distributions we used to generate the measurement data or our model. It is used in the Metropolis sampler to generate the next \(\mu\) value. This is sampled from the normal distribution, i.e., selected randomly.
# new µ-value sampled from normal distribution with fixed standard deviation
mu_proposal = norm(mu_current, proposal_width).rvs() The two corresponding distributions of the likelihoods and the prior are then calculated for the new (mu_proposal) and the old (mu_current) \(\mu\) value. Note that the corresponding \(\mu\) value is included in the calculation of the prior, i.e., if a new \(\mu\) value has been determined, then this must be used to determine the corresponding prior.
Finally, the probability distributions of the numerator of the Bayes' theorem are determined.
# for “old” and “new” mu-value calculate …
# … likelihoods
likelihood_current = norm(mu_current, 1).pdf(data).prod()
likelihood_proposal = norm(mu_proposal, 1).pdf(data).prod()
# … prior probabilities
prior_current = norm(mu_prior_mu, mu_prior_sd).pdf(mu_current)
prior_proposal = norm(mu_prior_mu, mu_prior_sd).pdf(mu_proposal)
# ….nominator of Bayes formula
p_current = likelihood_current * prior_current
p_proposal = likelihood_proposal * prior_proposalIf this piece of code is executed repeatedly, it is a simple hill climbing algorithm. After each run, the new \(\mu\) value becomes the old \(\mu\) value and is used to sample a new \(\mu\) value for which the probability distribution of the new numerator of the Bayes' theorem is determined (the new value from the previous run becomes the new old value). Changes in the \(\mu\) value are possible in both directions (the new value can be greater or smaller than the previous value). However, since we “sample” the new values from a normal distribution, the value will approach \(\mu = 0 \) in the long run.
However, we are not only interested in the final mean value of \(\mu\), but we also want to determine its probability distribution. And this is where the actual “trick” of MCMC comes in: We determine an acceptance probability from the two denominator probabilities
p_accept = p_proposal / p_current # acceptance probabilityand distinguish between two cases: if the probability of acceptance is
- greater than 1, then the new \(\mu\) value is accepted
- less than 1, then the new \(\mu\) value is accepted with a probability determined by the probability of acceptance.
And that's the complete MCMC algorithm! Surprisingly simple for something that looks extremely complicated at first glance, right?
Why did we choose these two cases in particular?
If we only accepted results with an acceptance probability greater than or equal to 1, the algorithm would converge towards the mean value \(\mu\) we are looking for. However, we are not only interested in this mean value; we also want to determine the probability distribution of \(\mu\). Therefore, we must occasionally accept acceptance probabilities that are less than 1. The probability with which we accept these is determined by their value. If the value of the acceptance probability is 0.3, for example, this means there is a 30 percent probability that we will accept this value for \(\mu\).
accept = np.random.rand() < p_accept # sample random number between 0 and 1 and compare to acceptance value
if accept:
cur_pos = proposal # Update value (set new value)And that's the entire MCMC algorithm! Amazingly simple for something that looks extremely complicated at first glance, right?
Note:
Why did we choose this example? The answer is relatively simple. The prior is a normal distribution, and the conjugate of a normal distribution is again a normal distribution, i.e., this example can also be calculated by hand, which makes it easier for us to verify the result obtained.
Finally, here is the complete Python code for you to try out for yourself (like most of the above description, it was taken from Thomas Wiecki's excellent blog post).
def sampler(data, samples=4, mu_init=.5, proposal_width=.5, plot=False, mu_prior_mu=0, mu_prior_sd=1.):
mu_current = mu_init
posterior = [mu_current]
for i in range(samples):
# suggest new position
mu_proposal = norm(mu_current, proposal_width).rvs()
# Compute likelihood by multiplying probabilities of each data point
likelihood_current = norm(mu_current, 1).pdf(data).prod()
likelihood_proposal = norm(mu_proposal, 1).pdf(data).prod()
# Compute prior probability of current and proposed mu
prior_current = norm(mu_prior_mu, mu_prior_sd).pdf(mu_current)
prior_proposal = norm(mu_prior_mu, mu_prior_sd).pdf(mu_proposal)
p_current = likelihood_current * prior_current
p_proposal = likelihood_proposal * prior_proposal
# Accept proposal?
p_accept = p_proposal / p_current
# Usually would include prior probability, which we neglect here for simplicity
accept = np.random.rand() < p_accept
if plot:
plot_proposal(mu_current, mu_proposal, mu_prior_mu, mu_prior_sd, data, accept, posterior, i)
if accept:
# Update position
mu_current = mu_proposal
posterior.append(mu_current)
return np.array(posterior)
# Function to display
def plot_proposal(mu_current, mu_proposal, mu_prior_mu, mu_prior_sd, data, accepted, trace, i):
from copy import copy
trace = copy(trace)
fig, (ax1, ax2, ax3, ax4) = plt.subplots(ncols=4, figsize=(16, 4))
fig.suptitle('Iteration %i' % (i + 1))
x = np.linspace(-3, 3, 5000)
color = 'g' if accepted else 'r'
# Plot prior
prior_current = norm(mu_prior_mu, mu_prior_sd).pdf(mu_current)
prior_proposal = norm(mu_prior_mu, mu_prior_sd).pdf(mu_proposal)
prior = norm(mu_prior_mu, mu_prior_sd).pdf(x)
ax1.plot(x, prior)
ax1.plot([mu_current] * 2, [0, prior_current], marker='o', color='b')
ax1.plot([mu_proposal] * 2, [0, prior_proposal], marker='o', color=color)
ax1.annotate("", xy=(mu_proposal, 0.2), xytext=(mu_current, 0.2),
arrowprops=dict(arrowstyle="->", lw=2.))
ax1.set(ylabel='Probability Density', title='current: prior(mu=%.2f) = %.2f\nproposal: prior(mu=%.2f) = %.2f' % (mu_current, pri-or_current, mu_proposal, prior_proposal))
# Likelihood
likelihood_current = norm(mu_current, 1).pdf(data).prod()
likelihood_proposal = norm(mu_proposal, 1).pdf(data).prod()
y = norm(loc=mu_proposal, scale=1).pdf(x)
sns.distplot(data, kde=False, norm_hist=True, ax=ax2)
ax2.plot(x, y, color=color)
ax2.axvline(mu_current, color='b', linestyle='--', label='mu_current')
ax2.axvline(mu_proposal, color=color, linestyle='--', label='mu_proposal')
#ax2.title('Proposal {}'.format('accepted' if accepted else 'rejected'))
ax2.annotate("", xy=(mu_proposal, 0.2), xytext=(mu_current, 0.2),
arrowprops=dict(arrowstyle="->", lw=2.))
ax2.set(title='likelihood(mu=%.2f) = %.2f\nlikelihood(mu=%.2f) = %.2f' % (mu_current, 1e14*likelihood_current, mu_proposal, 1e14*likelihood_proposal))
# Posterior
posterior_analytical = calc_posterior_analytical(data, x, mu_prior_mu, mu_prior_sd)
ax3.plot(x, posterior_analytical)
posterior_current = calc_posterior_analytical(data, mu_current, mu_prior_mu, mu_prior_sd)
posterior_proposal = calc_posterior_analytical(data, mu_proposal, mu_prior_mu, mu_prior_sd)
ax3.plot([mu_current] * 2, [0, posterior_current], marker='o', color='b')
ax3.plot([mu_proposal] * 2, [0, posterior_proposal], marker='o', color=color)
ax3.annotate("", xy=(mu_proposal, 0.2), xytext=(mu_current, 0.2),
arrowprops=dict(arrowstyle="->", lw=2.))
#x3.set(title=r'prior x likelihood \(\propto\) posterior')
ax3.set(title='posterior(mu=%.2f) = %.5f\nposterior(mu=%.2f) = %.5f' % (mu_current, posterior_current, mu_proposal, posteri-or_proposal))
if accepted:
trace.append(mu_proposal)
else:
trace.append(mu_current)
ax4.plot(trace)
ax4.set(xlabel='iteration', ylabel='mu', title='trace')
plt.tight_layout()
#plt.legend()
To start the code, we need two more lines.
np.random.seed(123)
sampler(data, samples=8, mu_init=-1., plot=True);In the first line, the random generator is initialized, which generates a sequence of pseudo-random numbers using the specified start value (here: 123). By specifying the start value in this way, the same sequence of pseudo-random numbers is obtained each time the program is started, thus ensuring a reproducible result. In the second line, the actual algorithm for the "measurement data" (data) determined from a normal distribution is initialized with eight runs and the value -1 as the initial value for the mean \(/mu\).
The following figures show an example of the result of executing the above code. For each iteration step, the following are displayed (from left to right)
- the prior probability distribution (i.e., our assumption about the distribution of \(\mu\) before we took the (measurement) data into account) with the red and blue vertical lines for the current and proposed new \(\mu\) value, respectively,
- the likelihood distribution as a green or red line, depending on whether the value for \(\mu\) is accepted or not, as well as the histogram of the data used in the background,
- the normalized posterior distribution, and
- the posterior samples (i.e. the \(\mu\)-values) that have been determined in all iterations so far
for a total of eight iteration steps.
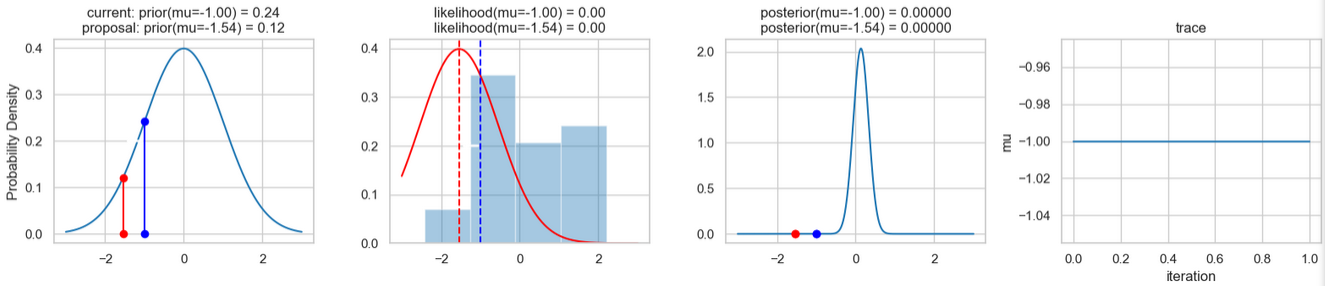
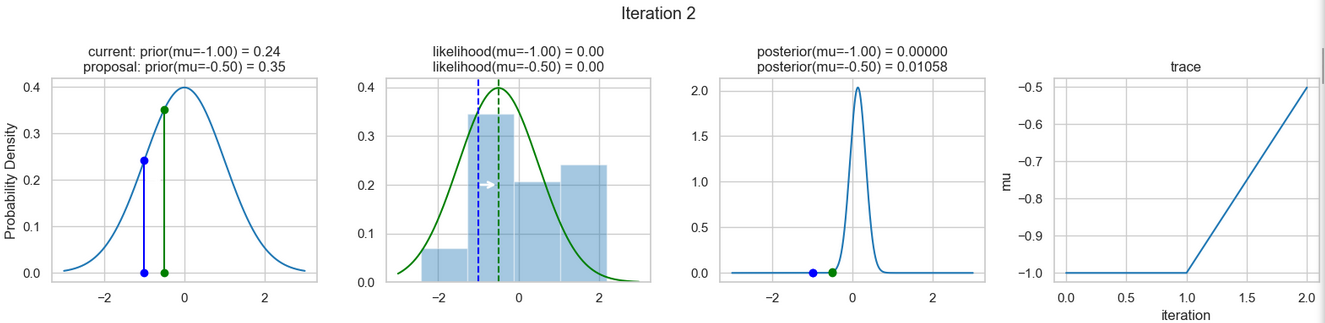
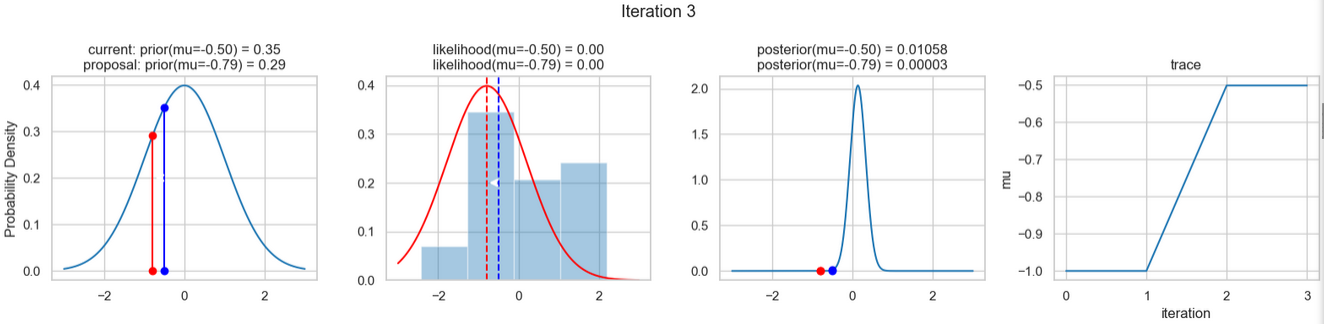
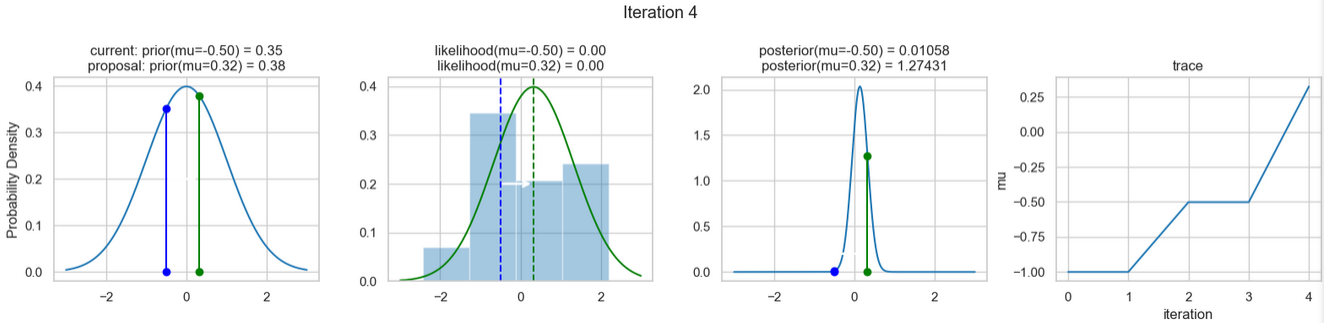
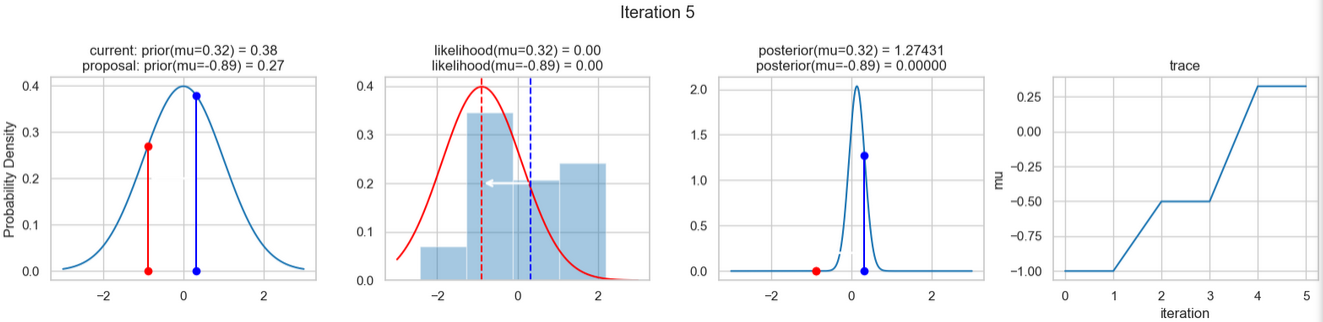
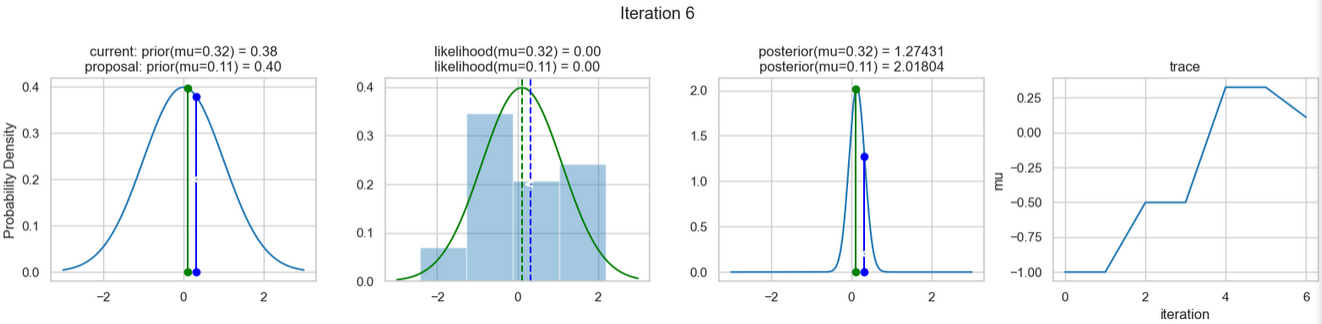
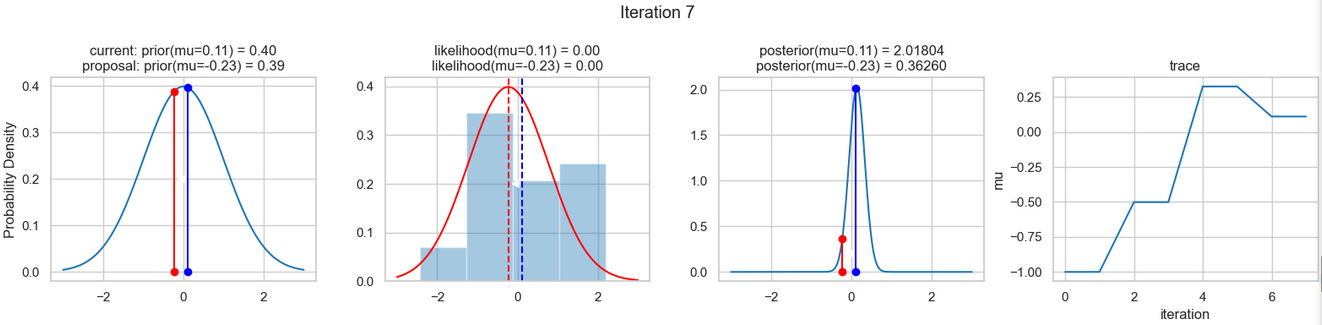
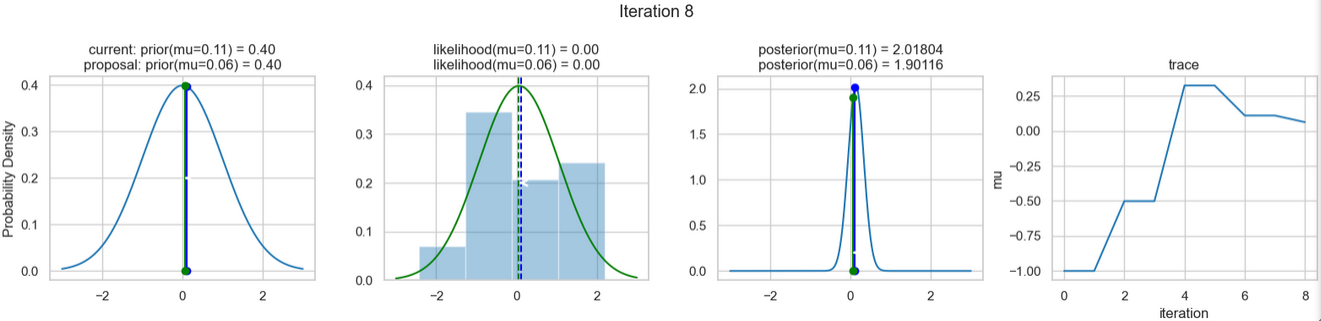
If you follow the results of the individual iterations, you will see that, over the course of the iterations, the µ value approaches the actual mean value \(\mu=0\) (i.e., the mean value underlying our data) (1st column), even if some of the calculated \(\mu\) values are discarded (red dots). Accordingly, the data is described better and better by our model (likelihood) (2nd column).
So far, so good! Now we only need a few more steps to complete this simple example.
Each of these iterations is based on the posterior probability distribution of our model! If we now perform a high number of iterations, we can obtain an approximation of the posterior probability distribution we are looking for (i.e., the application of Bayes' theorem).
Here is the corresponding Python code:
posterior = sampler(data, samples=15000, mu_init=1.)
fig, ax = plt.subplots()
ax.plot(posterior)
_ = ax.set(xlabel='sample', ylabel='mu');
The figure shows the result, which is called a trace and was executed for 15,000 samples (if you want to reproduce it on your computer: the calculation takes some time, so don't get impatient).
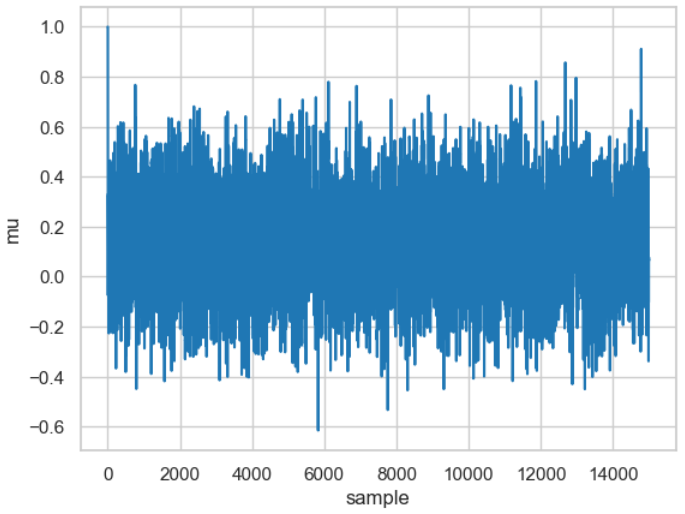
Result of the 15,000 calculated \(\mu\) values. The calculated \(\mu\) value is shown for each of the 15,000 samples. These vary around the actual mean (\(\mu=0\)).
From the trace, an approximation of the posterior probability distribution can now easily be determined by creating a histogram for it.
Here is the code used for this:
ax = plt.subplot()
sns.distplot(posterior[500:], ax=ax, label='estimated posterior')
x = np.linspace(-.5, .5, 500)
post = calc_posterior_analytical(data, x, 0, 1)
ax.plot(x, post, 'g', label='analytic posterior')
_ = ax.set(xlabel='mu', ylabel='belief');
ax.legend();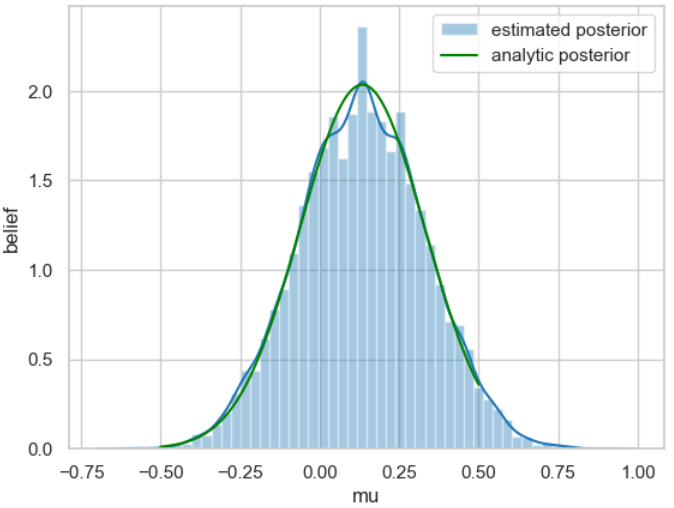
For better comparison, the posterior probability distribution calculated analytically at the beginning of the example is shown in green.
We are now essentially finished. We have taken a closer look at a method that allows us to use Bayes' theorem to determine the probability distribution. For the sake of completeness, we would like to address one further point at this point: the choice of the width of the normal distribution used for sampling.
There are two conflicting tendencies here. On the one hand, the width should not be too narrow, as sampling becomes extremely inefficient because it takes a long time to explore the entire parameter space. The curve then corresponds to a typical random walk, as shown by the result of the following code snippet.
posterior_small = sampler(data, samples=5000, mu_init=1., proposal_width=.01)
fig, ax = plt.subplots()
ax.plot(posterior_small);
_ = ax.set(xlabel='sample', ylabel='mu');
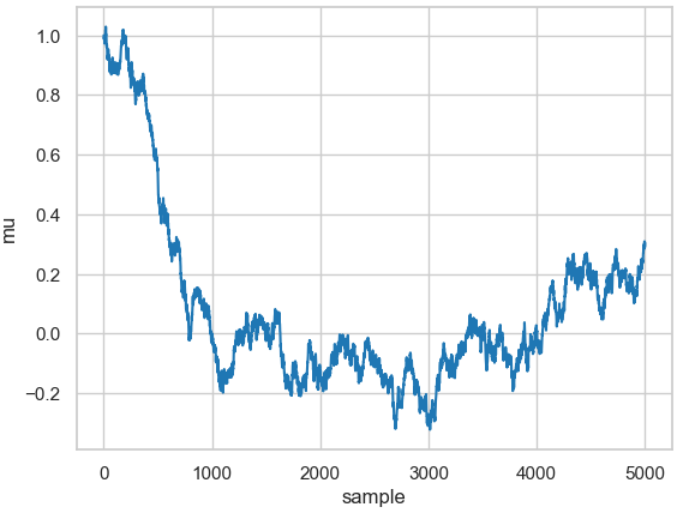
Result of a random walk for a relatively small value for the width (proposal_width=0.01).
On the other hand, choosing a too wide distribution can result in virtually no iteration leading to an accepted new value, as shown in the following code snippet.
posterior_large = sampler(data, samples=5000, mu_init=1., proposal_width=3.)
fig, ax = plt.subplots()
ax.plot(posterior_large); plt.xlabel('sample'); plt.ylabel('mu');
_ = ax.set(xlabel='sample', ylabel='mu');
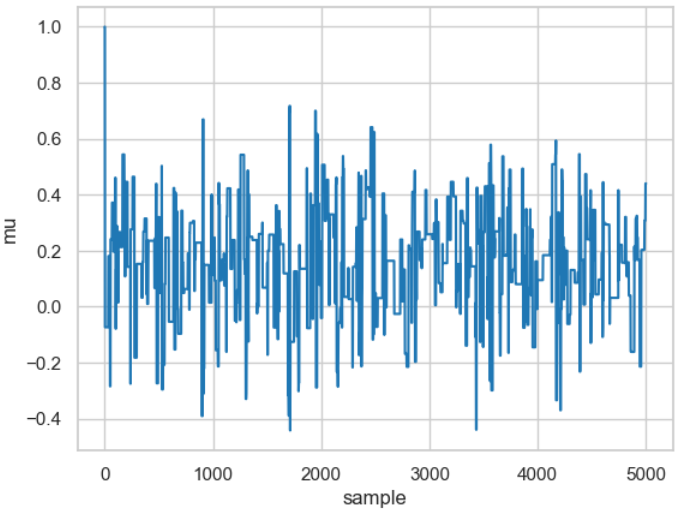
Result of a random walk for a relatively large value for the width (proposal_width=3).
The results for the two widths are based on the same posterior probability distribution, as shown below.
sns.distplot(posterior_small[1000:], label='Small step size')
sns.distplot(posterior_large[1000:], label='Large step size');
_ = plt.legend();
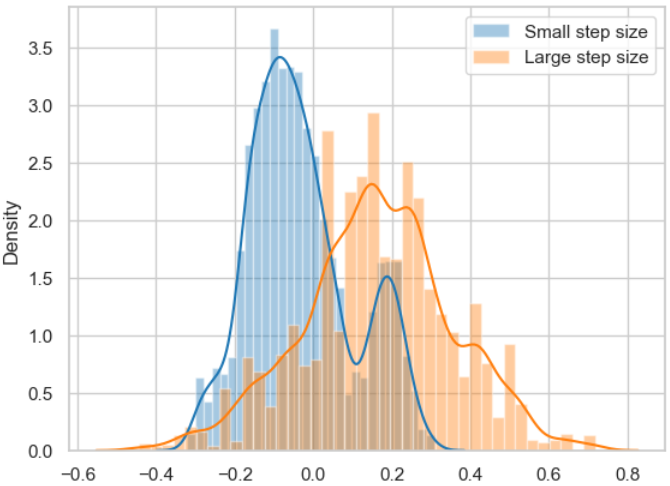
Posterior probability distributions for the two random walks determined above with small step size and large step size.
The figure clearly shows that the respective samples are not independent of each other, which is a key element for the successful application of the MCMC algorithm.
This naturally raises the question of how to determine whether the samples for the selected settings (in this case, the standard deviation of the normal distribution) are independent of each other and therefore efficient.
A commonly used metric for this is autocorrelation, which determines how correlated a sample is with all other samples.
Here is a small Python code with a result plot:
from pymc3.stats import autocorr
lags = np.arange(1, 100)
fig, ax = plt.subplots()
ax.plot(lags, [autocorr(posterior_large, l) for l in lags], label='large step size')
ax.plot(lags, [autocorr(posterior_small, l) for l in lags], label='small step size')
ax.plot(lags, [autocorr(posterior, l) for l in lags], label='medium step size')
ax.legend(loc=0)
_ = ax.set(xlabel='lag', ylabel='autocorrelation', ylim=(-.1, 1))
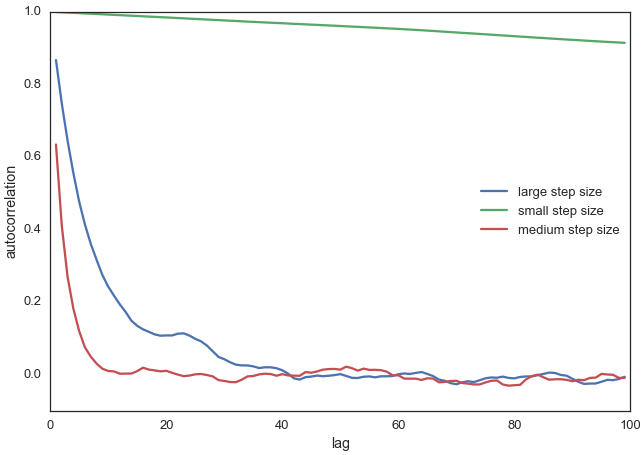
Autocorrelation plots (ACF) of random walks for different widths (step size). The x-axis (lag) shows the delay, i.e., the shift between the data sets. A lag of 1 compares each value with its immediate predecessor. The y-axis (autocorrelation) indicates the correlation value, which typically starts with a value of 1 for lag 0 (comparison with itself). A value of 1 means perfect positive correlation, -1 perfect negative correlation, and 0 no linear correlation.
It is clear that our original choice of width (proposal_width = 0.5, medium step size, red curve) shows significantly fewer correlations than in the other two cases (green and blue curves).
In practice, the best step size is determined automatically, for example, by the criterion that approximately 50% of the proposals are rejected.
But now we have finally reached the end of this example. We should now have a general understanding of the basic application of MCMC sampling to determine the posterior probability distribution based on existing measurement data, a model underlying the generation of the measurement data, and existing a priori knowledge.
As mentioned, this is a very simple example. In practice, much more complex steps are required—but the above principles usually remain valid. We will take a closer look at this in the following examples.


